SIRT1/PARP1 crosstalk: connecting DNA damage and metabolism
Map description below map
The best way to view the map is with PathVisio with the MIM plugin which requires 2 steps.
- Download the latest PathVisio-MIM installer if you have not already done so. To read more about PathVisio before you install it click here
- Download this map. It will be downloaded as a gpml file.
How to view the map using PathVisio.
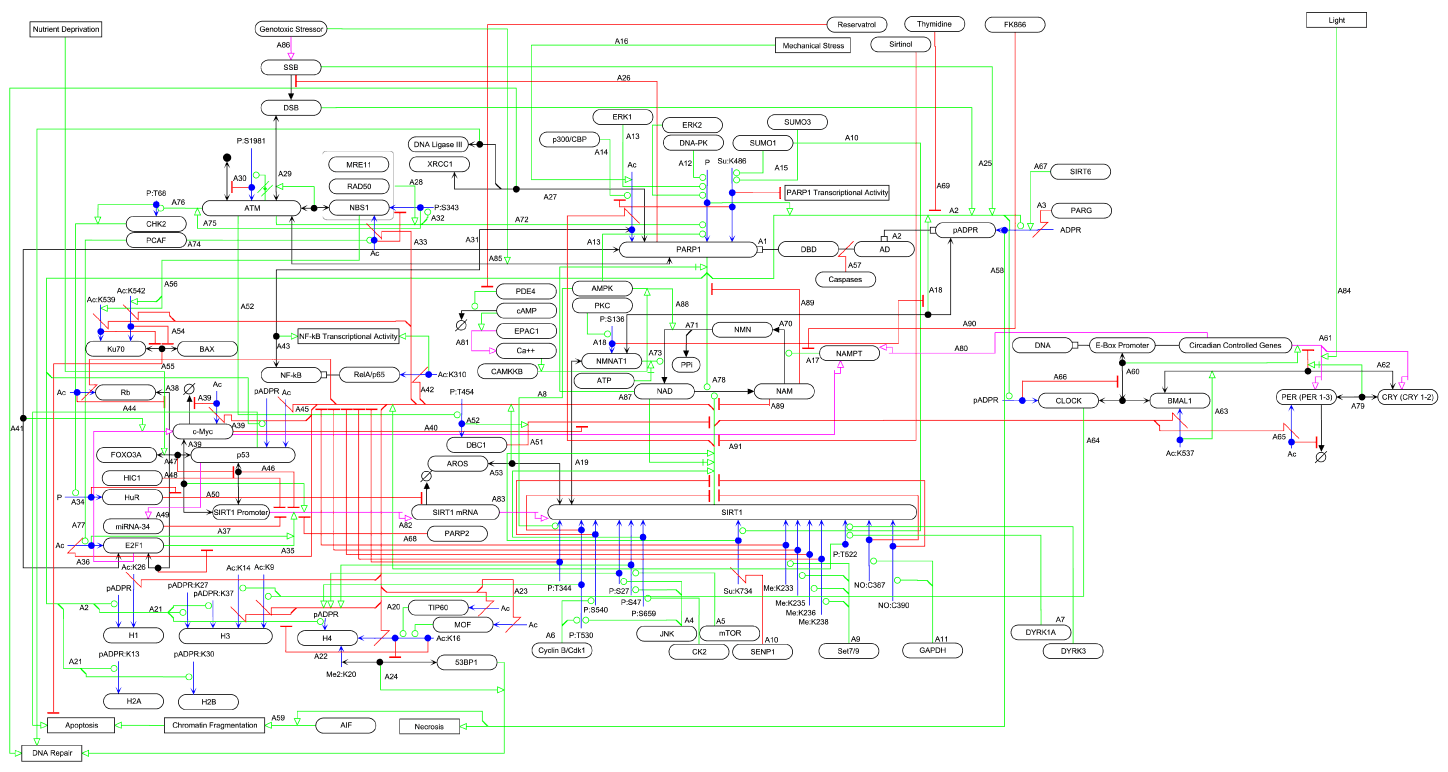
Augustin Luna, Mirit I. Aladjem and Kurt W Kohn
Implemented by : Margot Sunshine
An intricate network regulates the activities of SIRT1 and PARP1 proteins and continues to be uncovered.
Both SIRT1 and PARP1 share a common co-factor nicotinamide adenine dinucleotide (NAD+) and several common substrates,
including regulators of DNA damage response and circadian rhythms.
We review this complex network using an interactive Molecular Interaction Map (MIM) to explore the interplay between these two proteins.
Here we discuss how NAD + competition and post-transcriptional/translational feedback mechanisms create a regulatory
network sensitive to environmental cues, such as genotoxic stress and metabolic states, and examine the role of those
interactions in DNA repair and ultimately, cell fate decisions.
Link to paper
